In 902 CE, a fleet dispatched by the Umayyad Emirate of Córdoba arrived at Ibiza. The island was barely inhabited. Contemporary Andalusi writers rarely mentioned it at all. Whatever pre-conquest population existed had either fled or dwindled to near nothing. The newcomers — Imazighen clan groups, some Arabs, some Islamized Iberians — settled onto essentially empty land.
What happened next, genetically speaking, is what a new study published in Nature Communications1 sets out to reconstruct.
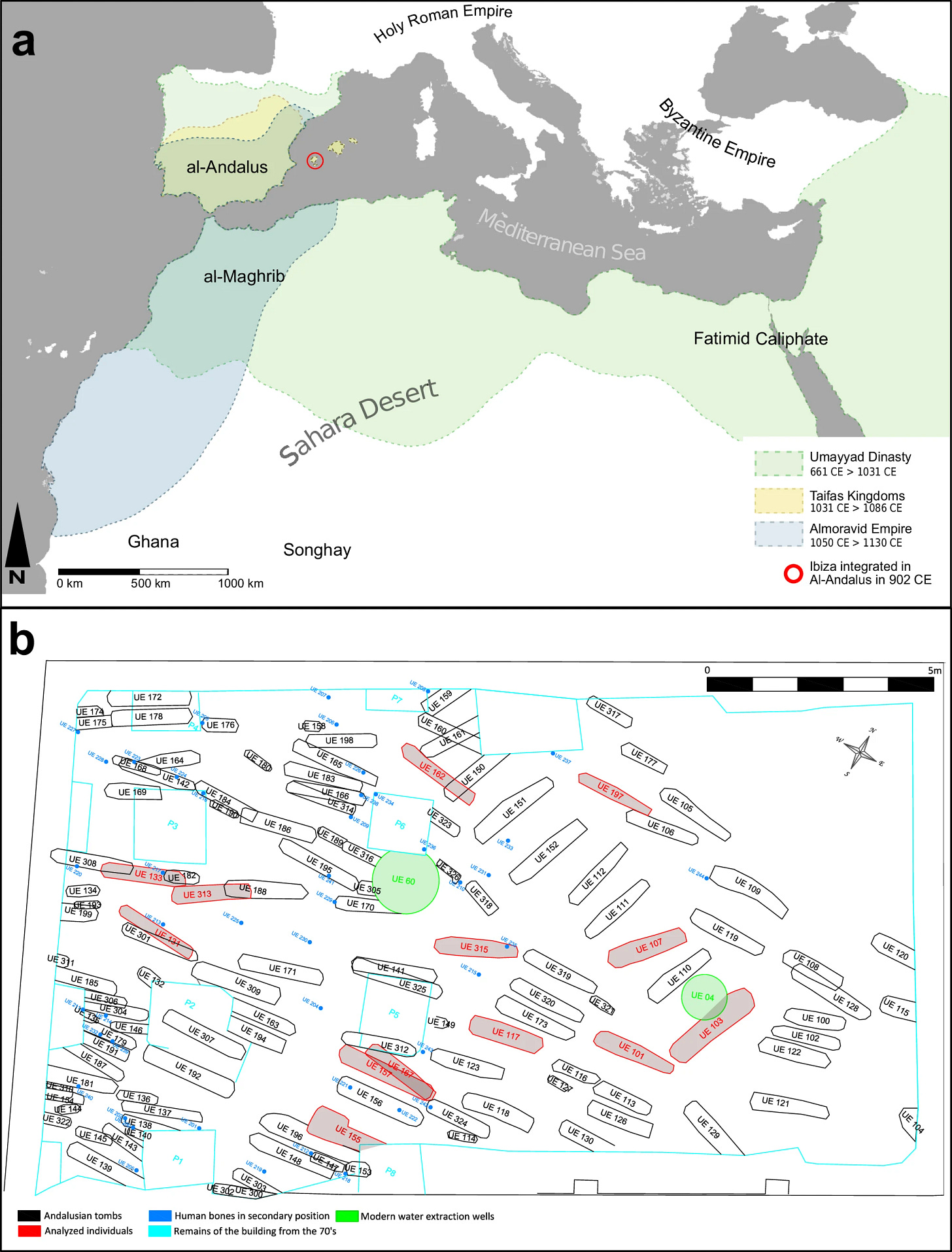
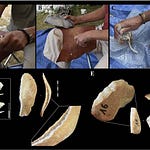
Researchers from the Centre for Palaeogenetics at Stockholm University analyzed ancient DNA recovered from 13 individuals buried in a section of the Maqbara of Madina Yabisa, the main urban Muslim cemetery of medieval Ibiza, discovered during construction work at 33 Bartomeu Vicent Ramon Street in Ibiza town. The burials date to 950–1150 CE. Simple earth pits, bodies placed on the right side facing southeast toward Mecca, no grave goods — with one exception, a burial containing two silver rings. These were people interred according to Islamic law, in a functioning urban cemetery on an island that had gone from empty to organized in a single generation.
From 30 sampled individuals, 13 yielded enough DNA for population genomic and metagenomic analyses. That’s a small number. What they show is not.
Who Was Buried Here
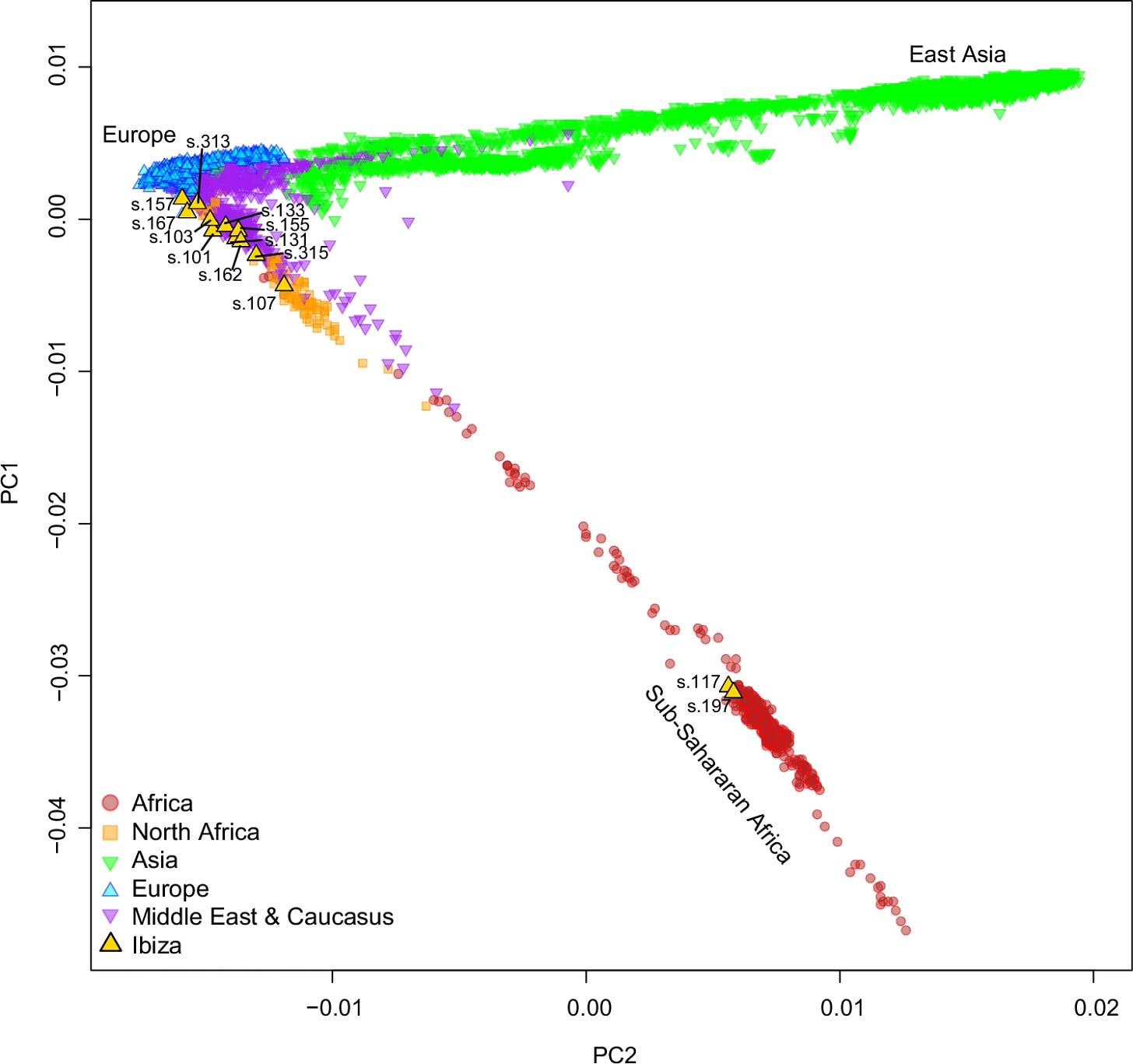
The genetic picture of these 13 individuals is anything but homogeneous. Principal component analysis places them across a wide swath of the ancestral landscape: two cluster within European population space, one sits close to North African populations, eight occupy positions somewhere between European, Middle Eastern, and North African, and two sit squarely in Sub-Saharan African genetic space.
The majority carry mixtures of Iberian and North African ancestry, consistent with the known demographic history of al-Andalus after the Islamic conquest of Iberia in 711 CE, when Imazighen (Amazigh, often called Berbers in older literature) formed the largest group of new arrivals. Y-chromosome haplogroups E1b1b1b1a1 and E1b1b1a1a1c2 — both common in North African Amazigh populations — appear in six of the nine males. One individual, s.107, clusters so closely with pre-European contact Canary Islanders and Moroccan Imazighen that the team interprets him as an Amazigh individual whose small European ancestry component likely reflects the ancient European-related ancestry already present in pre-Islamic North African populations, not recent admixture with local Iberians.
Two individuals, s.157 and s.313, are genetic outliers in the other direction: minimal North African ancestry, genomes that look much like pre-Islamic Iberians. Both are buried in a Muslim cemetery. The team reads them as muladíes — muwalladûn in Arabic, recently Islamized local Iberians who retained the genetic signature of their ancestry while adopting a new religious identity. The observation has a larger implication: on Ibiza in the 11th century, cultural and religious transformation was apparently decoupled from genetic admixture. People converted to Islam faster than they mixed.
Two People from Opposite Ends of the Sahel
The two Sub-Saharan individuals are the most striking members of this group.
Individual s.117 is male. His mitochondrial haplogroup is L3e1c, his Y-chromosome haplogroup E1b1a1a1a — both found primarily in Sub-Saharan African populations. When the team compared his genome against a comprehensive panel of modern African populations including groups from across Chad, Sudan, the Sahel, West Africa, and East Africa, he aligned most closely with present-day populations from southern Chad: the Sara and the Laal, speakers of a Nilo-Saharan language and an endangered language isolate, respectively.
Individual s.197 also carries Sub-Saharan ancestry — mitochondrial haplogroup L3b2 — but his genome tells a different story. He clusters with Gambian populations and the Senegal Bedik, groups from the Senegambia region of West Africa.
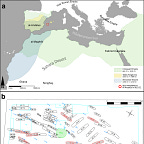
These two men, buried in the same small cemetery on a Mediterranean island, came from regions roughly 4,000 kilometers apart, connected by the trans-Saharan networks that Arabic sources describe in some detail. Historical records from Ibn ʿIdhārī, al-Bakrī, and the al-Bayān al-Mughrib tradition document the northward movement of enslaved individuals from the Lake Chad basin via the Kawar and Fezzan oases — through present-day Niger and Libya — toward North Africa and ultimately Iberia. Those same sources describe a military force of around 4,000 cavalry from the kingdom of Takrūr, in the middle-lower Senegal Valley, crossing into Iberia with Yūsuf b. Tāshfīn at the Battle of Zallaqa in 1086 CE. Takrūr had converted to Islam around 1030 CE, and trans-Saharan networks subsequently supplied the Almoravid armies with both captives and voluntary soldiers.
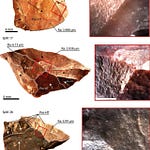
The radiocarbon dates of s.117 and s.197, corrected for marine reservoir effects given Ibiza’s island context, place both individuals after 1115 CE — in the second demographic pulse that reached the Balearics following the Almoravid conquest of Mallorca in 1115–1116 CE. The genomic evidence and the historical record align on the same point: these were people moved by the Almoravid military and slave systems, arriving on Ibiza as part of a larger resettlement and garrisoning process.
The coexistence of Chadian and Senegambian ancestry in a single small cemetery shows something the historical texts suggest but rarely make vivid: the al-Andalus of the 11th and 12th centuries was not drawing from one part of sub-Saharan Africa. It was drawing from both the western and central Sahel simultaneously, through distinct but overlapping networks.
Timing the Mixture
Beyond identifying ancestral origins, the team used haplotype-based local ancestry analysis to estimate when North African admixture actually occurred in the ancestors of these individuals.
The method works by measuring the length of uninterrupted chromosomal segments of North African ancestry. When two populations mix, the resulting chromosomes carry long stretches of one ancestry type unbroken by recombination. Each subsequent generation, recombination shuffles those stretches shorter. The rate of that decay maps onto generational time.
The population-level estimate, combining seven individuals radiocarbon-dated between 1073 and 1094 CE, places the main admixture event approximately 7.84 generations before 1080 CE. Using a generation time of 26.9 years, that translates to roughly 869 CE — predating Ibiza’s conquest in 902 CE by about 33 years, and likely reflecting admixture that began in mainland al-Andalus before or during the initial colonization. Individual-level estimates range from 2.49 to 7.81 generations, pointing to ongoing gene flow rather than a single founding event.
One individual, s.157 — one of the two with predominantly European ancestry — shows an admixture date going back approximately 16 generations, to around 519 CE. This is pre-Islamic. The team interprets this as an ancient, low-level North African genetic contribution predating the Muslim conquest, probably connected to the demographic transformations of the late Roman period. S.157 may represent a family line that had a small North African contribution centuries earlier, absorbed into the local Iberian population long before Ibiza became part of al-Andalus.
Two individuals, s.131 and s.315, show elevated runs of homozygosity consistent with consanguinity, with s.131’s pattern suggesting first-cousin parental relatedness. This kind of endogamy was documented among Imazighen communities and in other small medieval Iberian populations. S.131’s admixture timing of 3.47 generations suggests that both parents were already part of the admixed local gene pool of Ibiza — a community that had been mixing for generations and was now marrying within itself.
The Leprosy Case
Individual s.313 is one of the two with minimal North African ancestry, almost certainly a muladí. His bones show no diagnostic skeletal signs of leprosy — some facial elements are missing, which makes a definitive osteological assessment impossible, but what survives looks unremarkable.
His DNA tells a different story.
Metagenomic screening of s.313’s shotgun sequencing data detected Mycobacterium leprae, the bacterium responsible for leprosy. Target enrichment with a custom capture panel increased the read yield from roughly 29,000 reads to 128,800, enabling a mean genome coverage of 3.75x — enough for phylogenetic placement.
The M. leprae genome from s.313 belongs to genotype 2F, a clade containing seven ancient genomes from across medieval Europe, dated between approximately 650 and 1250 CE. The clade stretches from Hospital of Sant Llàtzer/Santa Margarida in Barcelona to Sigtuna, Sweden. Maximum parsimony analysis tentatively groups s.313 with an individual from medieval Denmark (Jorgen749, 1223–1279 CE), though this specific relationship was not strongly supported in the maximum likelihood reconstruction.
The distribution of genotype 2F across medieval Europe illustrates something that individual burial evidence tends to obscure: disease was moving across the continent along the same networks that moved people, goods, and armies. Ibiza was not isolated from that. A bacterium recovered from a Muslim muladí in the western Mediterranean shares a lineage with individuals buried in Sweden.
S.313’s burial was indistinguishable from others in the cemetery. No sign of marginalization, no separation from the community. This is consistent with both Islamic theological and legal frameworks of the period, which placed considerable emphasis on the obligation to care for the sick, and with a broader pattern, also documented in contemporary Christian cemeteries, of including individuals with leprosy in standard community burials rather than excluding them. Whether s.313 had visible disease at the time of death, or died before symptoms appeared, or had a subclinical infection that never progressed, cannot be determined from the available evidence.
Pathogens Across the Group
The metagenomic analysis found traces of multiple pathogens beyond M. leprae. Hepatitis B virus was detected in individual s.167 with high confidence, and lower-coverage signals were validated in three others; genotyping recovered genotypes A and D, both known from medieval European HBV diversity. Human parvovirus B19, the cause of fifth disease, was recovered from four individuals as partial genomes clustering within genotype 2, with trace-level signals in five more. The parvovirus genomes help fill a temporal gap in the European B19V record between roughly 950 CE and the 20th century, and their assignment to genotype 2 extends the known persistence of that lineage.
In at least 3 of the 13 individuals analyzed, the team identified molecular evidence of infection by primary pathogens. Including lower-coverage HBV signals brings that number to 5, or approximately 38% — comparable to rates reported from broad-spectrum pathogen screening at other medieval sites. Including Streptococcus pneumoniae (which can be carried asymptomatically) and a periodontal pathobiont, Parvimonas micra, detected in individual s.133, puts the count at 7 individuals, or about 54%.
It’s a small community, and more than half of the people whose DNA could be recovered were carrying some detectable pathogen.
What the Island Was
Ibiza before 902 CE barely existed in the written record. By the time these 13 people were buried — between 950 and 1150 CE — the island had become a node in a network that extended south to the Sahel, east to the Levant, and north into a Mediterranean world increasingly structured by the confrontation between Islamic and Christian polities.
The genomes reflect this. They show an initial colonization wave from mainland al-Andalus, dominated by Imazighen with some Arab and Islamized Iberian participants, followed by a second pulse after the Almoravid conquest of Mallorca in 1115–1116 CE that brought new arrivals from North Africa, the Iberian peninsula, and — demonstrably — the far side of the Sahara. The timing estimates suggest admixture that had been ongoing for generations by the time these individuals were alive, some of it recent enough that the chromosomal blocks of North African ancestry were still long and largely intact.
The two individuals with the deepest European ancestry — genetically local, culturally Muslim — had apparently converted without marrying into the arriving populations. Or at least their ancestors had. And one of them, without any visible sign of disease, was carrying Mycobacterium leprae genotype 2F, a lineage traveling from the Mediterranean to Scandinavia and back again through the bodies of medieval Europeans who left few records other than this.
Further Reading
Olalde, I. et al. “The genomic history of the Iberian Peninsula over the past 8000 years.” Science 363, 1237–1241 (2019).
Oteo-Garcia, G. et al. “Medieval genomes from eastern Iberia illuminate the role of Morisco mass deportations in dismantling a long-standing genetic bridge with North Africa.” Genome Biology 26, 108 (2025).
Pfrengle, S. et al. “Mycobacterium leprae diversity and population dynamics in medieval Europe from novel ancient genomes.” BMC Biology 19, 1–18 (2021).
Schuenemann, V.J. et al. “Ancient genomes reveal a high diversity of Mycobacterium leprae in medieval Europe.” PLOS Pathogens 14, 1006997 (2018).
Mühlemann, B. et al. “Ancient human parvovirus B19 in Eurasia reveals its long-term association with humans.” PNAS115, 7557–7562 (2018).
Kocher, A. et al. “Ten millennia of hepatitis B virus evolution.” Science 374, 182–188 (2021).
Fortes-Lima, C.A. et al. “Population history and admixture of the Fulani people from the Sahel.” American Journal of Human Genetics 112, 261–275 (2025).
Dury, G. et al. “The Islamic cemetery at 33 Bartomeu Vicent Ramon, Ibiza: investigating diet and mobility through light stable isotopes in bone collagen and tooth enamel.” Archaeological and Anthropological Sciences 11, 3913–3930 (2019).
Bennison, A.K. The Almoravid and Almohad Empires. Edinburgh University Press, 2016.
Rodríguez-Varela, R. et al. “Five centuries of consanguinity, isolation, health, and conflict in Las Gobas: A Northern Medieval Iberian necropolis.” Science Advances 10 (2024).
Rodríguez-Varela, R., Pochon, Z., Mas-Sandoval, A., Yaka, R., Fortes-Lima, C.A., García Rubio, A., et al. “Analysis of medieval burials from Ibiza reveals genetic and pathogenic diversity during the Islamic period.” Nature Communications 17, 2703 (2026). https://doi.org/10.1038/s41467-026-70615-9